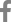Introduction
Rotavirus is the most common causal agent of acute diarrhea among children worldwide and is transmitted between individuals via the fecal-oral route1). Approximately half a million children under the age of five die from rotavirus infections every year2). Most of these children reside in low-income countries in Africa and Asia where access to safe water, sanitation, and medical care is often limited3).
Rotavirus, a member of the Reoviridae family, is classified into eight groups or species named A to H4). The viral genome consists of 12 segments of double-stranded RNA, which encode six structural proteins (VP1-VP4, VP6, and VP7) and six nonstructural proteins (NSP1-NSP6)5). The outer capsid comprises two proteins, the glycoprotein VP7 and the protease-sensitive VP4, which define the G and P genotypes, respectively, and relate to host specificity, virulence, and protective immunity6).
Rotavirus, when diagnosed in the clinical setting, cannot be distinguished from other forms of acute gastroenteritis caused by other microorganisms. Rotavirus infection can be accurately diagnosed by performing a rapid antigen detection of rotavirus in stool samples. Commercially available assays have been developed to detect rotaviruses in clinical samples, including electron microscopy (EM)7), virus culture8), enzyme immunoassay (EIA)9), latex agglutination (LA) assay10), polyacrylamide gel electrophoresis (PAGE)11), dot-blot hybridization assay12), and reverse transcription polymerase chain reaction (RT-PCR)13). Currently, the EIA and RT-PCR methods are the most popular assays for detecting, serotyping, and genotyping infectious rotavirus strains.
In South Korea, rotavirus is the most common cause of acute diarrhea in young children, particularly during the winter months14). Studies on the etiology of childhood diarrhea have shown that rotavirus is the most frequently diagnosed gastroenteritis agent detected in stool specimens obtained from Korean children hospitalized with acute diarrhea, accounting for 41.3-68% of all cases of gastroenteritis15-18).
The purpose of this study was to identify the distributions of G and P rotavirus serotypes on the basis of data collected on children hospitalized with rotavirus-related diarrhea in South Korea between 1989 and 2009. In addition, this review study discusses the genetic and antigenic diversity of Korean rotavirus strains that occur over two decades and which may indicate future trends of circulating rotavirus genotypes. The study data also address information about the control, estimation, and development of a nationwide rotavirus vaccine campaign in Korea.
Materials and methods
The study data were collected through two independent academic article searches of the MEDLINE database. The first search retrieved published studies from the PubMed-NCBI database (http://www.ncbi.nlm.nih.gov). This search used the terms "human," "rotavirus," and "Korea." The second search, which retrieved published articles from the KoreaMed database (http://koreamed.org/SearchBasic.php), used the terms "rotavirus" and "human." Both searches focused on hospitalized children and outpatients who had been hospitalized with acute gastroenteritis.
The genetic sequences of the G and P rotavirus genotypes were obtained from NCBI to create phylogenetic trees for analyzing the genetic diversity of the South Korean rotavirus strains (Supplementary Table 1). The phylogenetic trees were constructed using neighbor-joining algorithms19) from the Phylip package20). Evolutionary distances for the neighbor-joining analysis were based on the model described by Jukes and Cantor21). Tree topology was evaluated using the bootstrap resampling method with 1,000 replicates of the neighbor-joining dataset with the SEQBOOT and CONSENSE programs from the Phylip package.
Summary of rotavirus diagnostic assays
There are several differences in the clinical manifestations and epidemiologic patterns of rotaviral gastroenteritis and other forms of acute gastroenteritis22). However, a rotaviral infection is difficult to diagnose and cannot be identified solely on the basis of symptoms of gastroenteritis. An accurate rotavirus diagnosis requires routine laboratory confirmation such as the detection of virus or viral antigen or demonstration of a serologic response5). Currently, numerous assays have been developed to detect rotaviruses in clinical samples.
1. Electron microscopy
EM visualization is highly specific for rotavirus detection because these viruses have a distinctive morphology with a double-shelled capsid7,23-25). However, the experience and equipment required for an EM diagnosis are labor intensive, which can limit the routine detection of rotavirus for large numbers of clinical specimens. In addition, EM cannot distinguish between rotaviruses of different groups and serotypes.
3. Enzyme immunoassay
EIA is the preferred identification method in many laboratories because it is highly sensitive, specific, and adaptable to large sample volumes, and it does not require specialized equipment9,23,28). Additionally, EIAs allow VP6 subgroup or VP7 serotype determination because this technique uses serotype-specific monoclonal antibodies29,30).
5. Polyacrylamide gel electrophoresis
The PAGE technique is used to determine the electrophoretic patterns of the rotavirus dsRNA11,32). The viral dsRNA is first extracted and then analyzed by electrophoresis on a polyacrylamide gel, followed by silver staining. The silver staining technique allows the rotavirus to be identified on the basis of the dsRNA pattern, which can be displayed as either a "long" or a "short" electropherotype33).
6. Dot-blot hybridization assay
The dot-blot hybridization assay was developed on the basis of in situ hybridization of labeled rotavirus ssRNA transcripts. These transcripts adhere to heat-denatured rotavirus RNA immobilized on a nitrocellulose membrane. This dot-blot technique has been found to be 10- to 100-fold more sensitive than a confirmatory enzyme-linked immunosorbent assay (ELISA)12,34,35).
7. Reverse transcription polymerase chain reaction
RT-PCR was developed to improve identification of rotaviruses and is based on the amplification of rotavirus RNA13). This technique is theoretically much more sensitive than the ELISA, PAGE, and EM techniques for detecting rotaviruses in different clinical specimens as well as environmental samples36). RT-PCR has significantly reduced the incidence of rotavirus nontypeable strains, which are difficult to identify with less sensitive techniques. Recently, a combination of EIA and RT-PCR was used for detecting, serotyping, and genotyping the infectious rotavirus strains37).
Diagnostic assay detection of rotavirus in South Korea
The majority of previous studies in South Korea have used ELISA to determine rotavirus serotypes by using either VP6- or VP7-specific monoclonal antibodies for G1, 2, 3, and 4 serotypes14,16,38-40). However, the sensitivity and specificity of the ELISA tests are different between commercial ELISA kits. In recent years, with the application of RT-PCR, the incidence rates of nontypeable strains have drastically decreased. RT-PCR was initially used to determine the serotype of rotavirus after a positive ELISA test indicated rotavirus as the causal agent. In addition, previous studies have applied a PAGE assay technique, although this application was only used to characterize detected strains41-43).
Prevalence of G and P genotypes in South Korea
A study on the worldwide G and P rotavirus genotype distributions has indicated that at least 27 G and 35 P genotypes have been reported to date44). Epidemiological studies have shown that of these genotypes, five G (G1-G4, and G9) and three P (P[4], P[6], and P[8]) genotypes are the major genotypes associated with global human rotavirus infections45).
1. G1-G4 genotypes
G1, G2, G3, and G4 are major G genotypes that account for more than 93% of all human rotavirus infections in South Korea (Table 1, Fig. 1). The predominance of each genotype depends on the specific time period and geographic area of viral infections. Prior to 2000, G1 was the predominant genotype14,16,46,47), and studies have indicated that this genotype was predominant mostly between 2004 and 200948-51). The G2 and G3 genotypes were the predominant genotypes during the 2000-2001 and 2003-2005 rotavirus seasons, respectively43,52). Another study has indicated that the G4 genotype was predominant during the 2002-2003 and 2007-2008 seasons41,53). The incidence of the G2, G3, and G4 genotypes generally decreased after the introduction of rotavirus vaccines, whereas the incidence of the G1 genotype slightly increased (Fig. 1). Genetic analysis revealed that genetic variation occurred over time despite the lack of available gene sequence data for the specific South Korean genotypes41,42,54,55). Phylogenetic analysis of the G1 genotype indicated that the G1 strains were clustered in lineage IV, lineage II, and mostly in lineage I and displayed distinctions from the G1 Wa prototype strain, which belongs to lineage III (Fig. 2A)42,54,55). Thirteen available Korean G2 strains were analyzed during the 2000-2001 and 2007-2009 seasons47,54). According to the analyses, 10 strains were clustered into lineage V and three other strains were clustered into lineage IV (Fig. 2A). Genetic analysis of the G3 and G4 genotypes showed that these strains were conserved and clustered into lineages III and Ie, respectively (Fig. 2A)41,54).
2. P[8], P[4], and P[6] genotypes
P[8], P[4], and P[6] are the major P genotypes, and these strains account for approximately 99% of all human rotavirus infections occurring in South Korea (Table 1, Fig. 1). P[8] was the most prevalent genotype, and studies report that this genotype constituted over one-half of all P genotypes identified in many parts of Korea during the study period46,48-53,56). P[6] and P[4] were the second and third most common P genotypes detected, respectively. Similarly, some additional studies show that the P[6] and P[4] genotypes were predominant during specific time periods and in particular geographic areas41,43,52,53). Phylogenetic analysis of the P[8] genotype revealed that these strains clustered into lineage IV and primarily lineage III (Fig. 2B)54,57). Phylogenetic analysis of the P[6] and P[4] genotypes revealed that these strains clustered into lineages V and I, respectively (Fig. 2B).
Clinical significance of unusual G genotypes in South Korea
1. G9 genotype
G9 is known to be the .fth most common genotype, after the G1-G4 genotypes, currently circulating in the human population. The first G9 rotavirus, WI61, was detected in children in the USA in 198358). After this point in time, G9 strains have been widely reported as the causative agents of diarrhea in children and have been recognized in many countries as one of the most widespread and emerging genotypes57). In South Korea, the first G9 strain was detected during the 2000-2001 rotavirus season43). Since then, G9 strains have been detected over consecutive seasons across the country (Table 1). The G9 strain was identified primarily in rural health care centers, and this genotype was found to be responsible for 7.4% to 39% of rural infections51-54,56,59); however, the rate of infection was much lower, only 1% to 3%, in urban hospitals41,48-50,60). G9 genotype detection generally improved after the introduction of rotavirus vaccines (Fig. 1). The low detection rate of G9 strains may be owing to the fact that the current vaccine formulations do not include the G9 antigen.
Currently, six lineages and 11 sublineages of the VP7 gene of G9 rotaviruses have been described in the research literature. A sequence analysis of the VP7 gene of various Korean G9 isolates demonstrated that the recent Korean G9 isolates belong to lineage III, within sublineage d, which also includes other contemporary G9 strains that are prevalent throughout the world (Fig. 2A). The recent increase in the incidence of global G9 human rotavirus infections indicates that type G9 has established itself as a globally common rotavirus G serotype of clinical importance45). Phylogenetic analysis of all 11 gene sequences of the three Korean G9P[8] strains revealed that these strains belonged to genogroup I of the Wa-like genotype constellation, which is one of the two main genotype constellations known to occur among epidemiologically important widespread human rotavirus A strains61).
2. G12 genotype
The first G12 rotavirus, L26, was detected in the Philippines in 19879). Currently, G12 genotypes are classified into four genetic lineages, I to IV62). In South Korea, the G12 genotype was first detected during the 2002-2003 season53). These strains were then intermittently detected in subsequent studies between 2004 and 200948,50,54). Sequence analyses of G12 genotypes in Korea are currently available for four strains (CAU195, CAU214, Seoul-553, and Seoul-681)48,54). These four Korean G12 strains are clustered into lineage III and show high sequence identity with strains isolated in Thailand, Germany, and the United States, which suggests that the Korean G12 strains are closely related to the recently reported G12 strains that emerged worldwide (Fig. 2A)54,63). Phylogenetic analysis of all 11 gene sequences revealed that these genotypes belonged to genogroup I of the Wa-like genotype constellation64). These results suggest that G12 genotypes not only are persistent but also may have adapted and evolved in the South Korean population.
3. G10 and G11 genotypes
Research analyses show that G10 and G11 rotavirus strains have rarely been detected in South Korea. Until now, only one G10 strain has been detected in Korea, and this occurred during the 2004-2005 season60). G11 strain analysis indicated that this strain was initially detected and characterized in the 2005-2006 season65) and was identified again during the 2008-2009 season49). However, additional studies are needed to investigate the importance and distribution of these rotavirus strains in the Korean population because only a few studies that specifically examine these G10 and G11 rotavirus strains have been conducted as yet (Table 1).
Clinical significance of unusual P genotypes in South Korea
1. P[9] genotype
The P[9] genotype is frequently detected in cats66), and the first P[9] human rotavirus, the K8 strain with G1 speci.city, was isolated from a 14-year-old schoolboy in Hokkaido, Japan in 197767). Although P[9] genotypes have been detected in most of the rotavirus surveillance studies in South Korea, there has been no significant emergence of P[9] genotypes in the Korean population43,48-51,53,59,60). Research has indicated that the majority of the P[9] strains carry G1, G2, and G3 specificity, except for a few isolates that were identified to have both G9 and G12 specificity48,51,53). Currently, no studies have focused on this genotype; therefore, further analysis of the circulating Korean P[9] genotypes is needed to understand their origin, genetic variation, and potential impact on the human population (Table 1).
2. P[3], P[10], and P[25] genotypes
To date, only one P[3] genotype and one P[25] genotype (unpublished data, KC140588) were detected during the 2003-2005 and 2011-2012 rotavirus seasons, respectively52). A total of five P[10] strains were detected in South Korea during three different rotavirus seasons43,51,60). However, the genetic profiles of the P[3] and P[10] genotypes have not been analyzed, even though the Korean P[25] genotype was found to be closely related to the P[25] genotypes isolated from Bangladesh (Dhaka6), Nepal (KTM368), and India (CRI10795) (unpublished data) (Fig. 2B). Because of these similarities and the implications for future rotavirus outbreaks, additional analysis of these genotypes is necessary (Table 1).
Prevalence of G and P genotypes and future implications for a nationwide rotavirus vaccine program in South Korea
Surveillance studies of the G and P genotypes in South Korea between 1989 and 2009 have provided important new information about rotavirus strain diversity. These studies have shown that there are six broad types of rotavirus strains, i.e., G1.G4, G9, and P[8]. These results are in agreement with the worldwide trend of the G and P rotavirus genotypes, in which a total of five G and P combinations (G1P[8], G2P[4], G3P[8], G4P[8], and G9P[8]) account for approximately 90% of all human rotavirus infections68).
Recently, two effective rotavirus vaccines have become available. The first vaccine, Rotarix (GlaxoSmithKline Biologicals, Rixensart, Belgium), is based on an attenuated G1P[8] rotavirus, whereas the other vaccine, RotaTeq (Merck & Co., Whitehouse Station, NJ, USA), is a human-bovine reassortant vaccine that contains the genotypes G1-G4 and P[8]. Rotarix has been approved in 90 countries worldwide and has shown an 80.5% efficacy against non-G1P[8] genotypes, including the G9P[8] genotype69-71). RotaTeq has been filed in more than 100 countries and has been reported to reduce hospital admission and emergency department visits by 94.5% after the third dose72). These two vaccines were introduced in South Korea in September 2007 and July 2008, respectively, and have been shown to be effective, safe, and economical through a nationwide rotavirus vaccination campaign73-75). In addition, the introduction of the RotaTeq vaccine in a three-dose regimen has reduced the rate of symptomatic rotavirus infections in South Korean infants by approximately 63.2% over a 5-year period73).
Conclusions
Rotavirus is the most common causal agent of acute diarrhea among children worldwide and is the most common causal agent of gastroenteritis in infants and young South Korean children. Currently, many assays are available for virus detection in clinical samples. The most common tests used for detection are the EIA and RT-PCR tests, which are used for detecting, serotyping, and genotyping virus infections. This review analyzed the epidemiology of G and P rotavirus genotypes on the basis of data between 1989 and 2009. All the available published articles on rotavirus studies in South Korea were retrieved from either the PubMed or KoreaMed databases.
The review analysis showed that the G genotypes G1-G4 were consistent and predominant, and that G9 strains were frequently detected. Other G genotypes such as G12, G11, and G10 were detected in some geographic settings; however, it was difficult to estimate the epidemiological importance of these genotypes owing to limited analyses (Table 1). With regard to the P genotypes, P[8] was found to be responsible for more than 53% of the infections, followed by P[6] and P[4] at 24% and 20%, respectively (Table 1). P[9] genotypes were reported in numerous studies, whereas P[3], P[10], and P[25] genotypes were rarely detected (Table 1). In general, the distributions of the G and P genotypes were found to be related to temporal and geographical fluctuations.
This study also addressed the application and effectiveness of rotavirus vaccines. However, further analysis is needed to determine the effectiveness and impact of these vaccines on the current and future trends of rotavirus genotypes. In addition, further characterization of the unexpected effects of rotavirus vaccines, such as vaccine-induced diseases, herd immunity, and changes in host susceptibilities, is also needed.





 PDF Links
PDF Links PubReader
PubReader PubMed
PubMed Download Citation
Download Citation